“Linking fast and slow: The case for generative models”
A pervasive challenge in neuroscience is testing whether neuronal connectivity changes over time due to specific causes, such as stimuli, events, or clinical interventions. Recent hardware innovations and falling data storage costs enable longer, more naturalistic neuronal recordings. The implicit opportunity for understanding the self-organised brain calls for new analysis methods that link temporal scales: from the order of milliseconds over which neuronal dynamics evolve, to the order of minutes, days, or even years over which experimental observations unfold. This review article demonstrates how hierarchical generative models and Bayesian inference help to characterise neuronal activity across different time scales.
Crucially, these methods go beyond describing statistical associations among observations and enable inference about underlying mechanisms. We offer an overview of fundamental concepts in state-space modeling and suggest a taxonomy for these methods. Additionally, we introduce key mathematical principles that underscore a separation of temporal scales, such as the slaving principle, and review Bayesian methods that are being used to test hypotheses about the brain with multiscale data. We hope that this review will serve as a useful primer for experimental and computational neuroscientists on the state of the art and current directions of travel in the complex systems modelling literature.
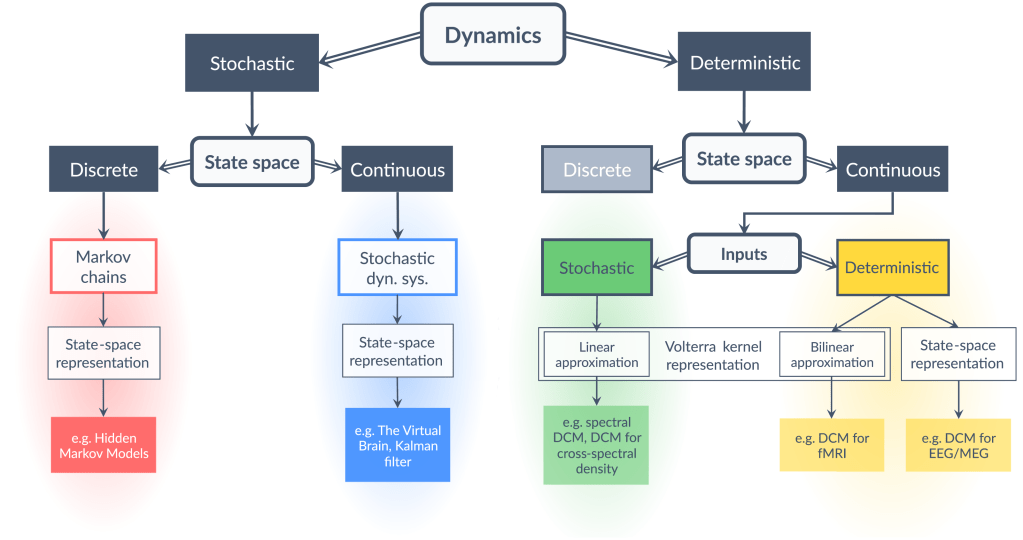
The key factors guiding the selection of a particular framework are the nature of dynamics (discrete or continuous) and the nature of state space (stochastic or deterministic). In addition, the nature of the inputs (stochastic or deterministic) has relevance for continuous deterministic systems. In the particular case of continuous deterministic systems with stochastic inputs, one can use a linear response function, which is the first-order term of the Volterra kernel representation of the system, to directly approximate the outputs from the inputs without reference to the states. Effectively, this implies that the dynamics do not need to be integrated over time, which greatly simplifies model inversion. In all other cases, model inversion entails tracking the states or their distribution through time.
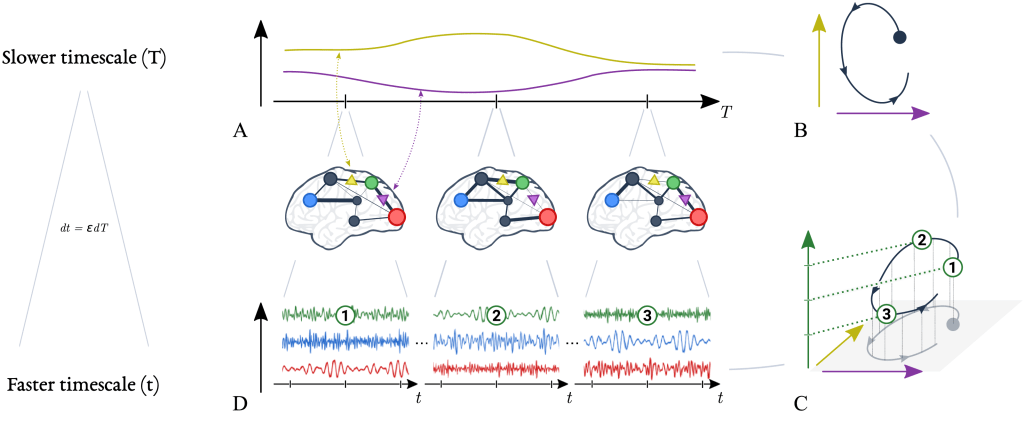
(A) Slow quantities in the brain, such as synaptic efficacy between regions, exhibit large time constants and evolve slowly over time. The evolution of two slow variables are illustrated here, as yellow and purple lines.
(B) The evolution of slow variables can also be represented by dynamics in a slow state space.
(C) Importantly, for every location in the slow state space (horizontal axes), there is a corresponding mode of fast dynamics (vertical axis). Three modes are depicted here, numbered 1–3. These fast dynamics give rise to rapid brain signals, such as field potentials in pyramidal neurons (D). The mathematical relationship between the slow and the fast timescale is given by dt = εdT (ε ≪ 1); in other words, the dynamics at faster scale t unfolds over a fraction (ε) of the slower scale T.
In summary, the brain is understood to navigate slowly (A) through a repertoire of fast stable dynamics (D). Crucially, the slow variables are directly linked to the dynamics of the fast variables (C). Similarly, changes in the fast variables’ dynamics can be attributed to changes in the slow variables. Therefore, modelling the complex dynamics of multiscale dynamical systems can be simplified by focusing on the dynamics of the slow variables and the mapping from slow to fast variables.
The mechanisms of interest for brain research generally span multiple temporal scales. Because of this apparent complexity, here we returned to first principles and pursued established mathematical paths. Evaluating hypotheses amounts to constructing generative models of our observations, for which there are readily available methods. The slaving principle means that slow and fast processes can be analysed separately: we can therefore model each timescale individually, lending a hierarchical structure to our generative models. Moreover, the slaving principle entails switching dynamics, for instance, of brain networks dynamics, which licenses the combination of discrete models (e.g., HMMs) for slow changes in brain states and continuous models of fast neuronal dynamics, as we have illustrated.
Evaluating and comparing the evidence for hypotheses regarding multiscale time series requires us to invert hierarchical temporal models, that is, to compute the posterior distribution over model parameters. In most cases, the inversion problem cannot be solved exactly because of both the complexity and hierarchical structure of the model. This forces us to approximate the posterior distribution, a task for which variational Bayes methods are preferred.
In particular, variational Bayes can be combined with empirical Bayes to invert arbitrary hierarchical models with empirical priors at intermediate levels. This suggests a simple procedure to invert hierarchical temporal models: first, model the fast temporal scale by fitting a model to each time window of time series data, and then invert these models to obtain a time series of estimated parameters (posteriors), which are expected to change slowly. Finally, use the time series of these slowly changing parameters as observations for the level above in the hierarchy.
In conclusion, by combining hierarchical state-space models with Bayesian analysis methods, time series data with different temporal scales can be linked and their interactions investigated. This has widespread application within neuroscience research, as well as in empirical science more generally.
