“From reductionism to realism: holistic mathematical modelling for complex biological systems”
At its core, the physics paradigm adopts a reductionist approach, aiming to understand fundamental phenomena by decomposing them into simpler, elementary processes.
While this strategy has been tremendously successful in physics, it has often fallen short in addressing fundamental questions in the biological sciences.
This arises from the inherent complexity of biological systems, characterized by heterogeneity, polyfunctionality and interactions across spatio-temporal scales.
Nevertheless, the traditional framework of complex systems modelling falls short, as its emphasis on broad theoretical principles has often failed to produce predictive, empirically grounded insights.
To advance towards actionable mathematical models in biology, we argue, using neuroscience as a case study, that it is necessary to move beyond reductionist approaches and instead embrace the complexity of biological systems—leveraging the growing availability of high-resolution data and advances in high-performance computing.
We advocate for a holistic mathematical modelling paradigm that harnesses rich representational structures such as annotated and multilayer networks, employs agent-based models and simulation-based approaches and focuses on the inverse problem of inferring system dynamics from observations.
We emphasize that this approach is fully compatible with the search for fundamental biophysical principles and highlight the potential it holds to drive progress in mathematical biology over the next two decades.
… biological systems, spanning from entire ecosystems to sub-cellular processes, have proved to be more impregnable to mathematical analysis. This is why it was first developed as a mostly descriptive science before moving to theoretical concerns.
While rich structures such as multilayer, annotated or hyper-networks offer the expressiveness necessary to model a range of interacting biological processes, one […] challenge is that many of these processes evolve over multiple time- and length-scales.
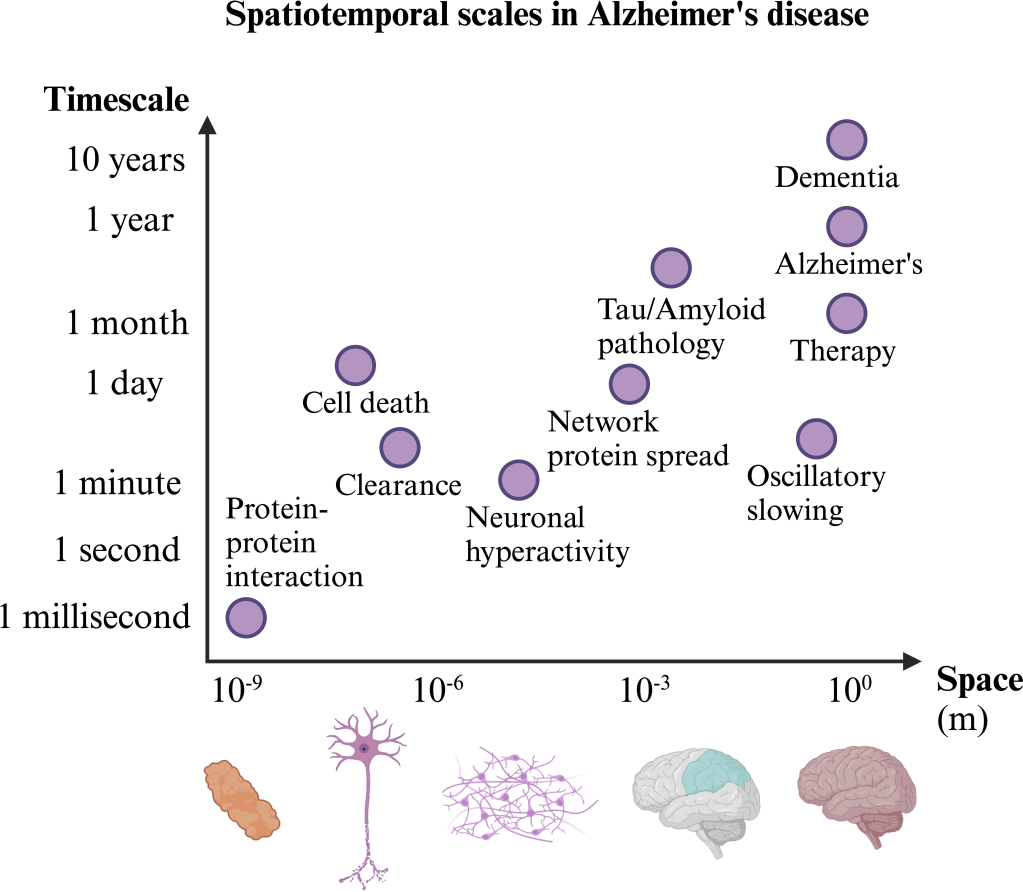
Complex biological processes, such as the development of neurodegenerative diseases such as AD, evolve over a range of interacting spatio-temporal scales. Focusing on AD, protein misfolding ultimately leads to the spread of toxic tau and amyloid-β proteins through axonal fibres, which is partially cleared through cell regulation. Ultimately, this leads to cell death and pathological neural dynamics. This manifests in tau and amyloid pathology and, finally, cognitive decline which can be combatted with therapy. The MM of this process must take into account the full range of spatio-temporal scales to build a mechanistic understanding of AD.
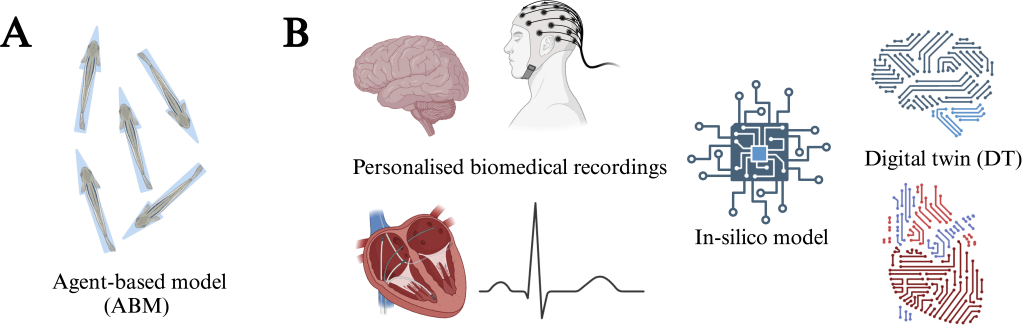
(A) ABMs (agent-based models )are powerful tools for describing the behaviour of heterogeneous populations of agentic units. Examples include the collective dynamics of swarms and flocks, where each agent can be modelled with its own dynamical properties.
(B) DT (digital twins) are simulation-based models of physical systems implemented in a computer. One application of biomedicine is to use personalized recordings of neural activity or heartbeat data to calibrate individual in silico models. Ultimately, this will lead to a virtual version of the biological system where interventions can be tested via simulation.
The traditional approach to modelling empirical observations of a dynamical system is to attempt to write down differential equations that describe the behaviour of the system.
Next, one attempts to find the parameters of this model that best explain the available data.
However, in large-scale complex systems, writing down a mechanistic model becomes a difficult challenge. For example, while Hodgkin and Huxley were able to isolate and study the dynamics of a single neuron in a model organism, the same cannot be performed for a macroscopic brain region in a human.
However, modellers now have access to an unprecedented amount of empirical data, i.e. neural recordings. This has inspired the development of a modern data-driven approach to dynamical systems, specifically the area of system identification. In particular, both parameter inference and system identification are examples of an inverse problem, which involves calculating a set of causal factors, be it parameter or model selection, from a set of observations.
Many methods for analysing nonlinear dynamics from high-dimensional data rely on the manifold hypothesis, which argues that high-dimensional data arising from complex systems are typically clustered around a lower-dimensional manifold.
As a result, the dynamics evolve in a latent space with fewer dimensions. This hypothesis justifies the use of dimensionality reduction methods such as principal component analysis, –distributed stochastic neighbour embedding and uniform manifold approximation and projection, which are ubiquitous in biological data analysis.
Such an assumption is also prevalent in data-driven approaches to dynamical systems, where much focus is placed on learning low-dimensional latent representations of high-dimensional dynamics.
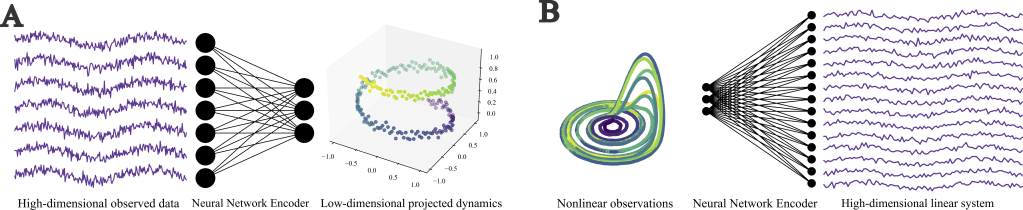
(A) High-dimensional data from complex biological systems is often clustered onto a low-dimensional manifold. Using data science techniques such as variational auto-encoders, we can discover the low-dimensional latent dynamics of the process.
(B) Following the theory of the Koopman operator, nonlinear dynamics can be ‘lifted’ into high-dimensional spaces where the dynamics are approximately linear. This projection can be achieved with deep learning techniques such as neural networks.
[…] Theoretical biology should turn towards complex, empirical models with predictive capabilities, even if these models occasionally complicate traditional mathematical analyses. Nevertheless, this shift does not negate the importance of identifying fundamental theoretical principles that govern biological systems, but instead, supports it.
Unlike physics, biology still lacks clear translational, foundational principles. Thus, the field urgently requires the development of a ‘theoretical physics for living systems’
